Small Large Language Model might sound like a bit of an oxymoron. However, I think it perfectly describes the class of LLMs in the 1-10 billion parameter range like Llama and Phi 3. In the last few days, Meta and Microsoft have both released these open(ish) models that can happily run on normal hardware. Both models perform surprisingly well for their size, competing with much larger models like GPT 3.5 and Mixtral. However, how well do they generalise to new unseen tasks? Can they do biology?
Introducing Llama and Phi
Meta's offering, Llama 3 8B, is an 8 billion parameter model that can be run on a modern laptop. It performs almost as well as Mixtral 8x22B mixture-of-expert model, a model 22x bigger and compute intensive.
Microsoft's model, Phi 3 mini, is around half the size of Llama 3 8B at 3.8 billion parameters. It is small enoughthat it can run on a high end smartphone at a reasonable speed. Incredibly, Phi actually beats Llama 3 8B, which is twice as big, at a few popular benchmarks including MMLU which approximately measures "how well does this model behave as a chatbot" and HumanEval which measures "how well can this model write code?".
I've also read a lot of anecdotal evidence about people chatting to these models and finding them quite engaging and useful chat partners (as opposed to previous generation small models). This seems to back up the benchmark performance and provide some validation of the models' utility.
Both Microsoft and Meta have stated that the key difference between these models and previous iterations of their smaller LLMs is the training regime. Interestingly, both companies applied very different training strategies . Meta trained Llama3 on over 15 trillion tokens (words) which is unusually large for a small model. Microsoft trained Phi on much smaller training sets curated for high quality.
Can Phi, Llama and other Small Models Do Biology?
Having a model small enough to run on your phone and generate funny poems or trivia questions is neat. However, for AI and NLP practitioners, a more interesting question is "do these models generalise well to new, unseen problems?"
I set out to gather some data about how well Phi and Llama 3 8B generalise to a less-well-known task. As it happened, I have recently been working with my friend Dan Duma on a test harness for BioAsq Task B. This is a less widely-known, niche NLP task in the bio-medical space. The model is fed a series of snippets from scientific papers and asked a question which it must answer correctly. There are four different formats of question which I'll explain below.
The 11th BioASQ Task B leaderboard is somewhat dominated by GPT-4 entrants with perfect scores at some of the sub-tasks. If you were somewhat cynical, you might consider this task "solved". However, we think it's an interesting arena for testing how well smaller models are catching up to big commercial offerings.
BioASQ B is primarily a reading comprehension task with a slightly niche subject-matter. The models under evaluation are unlikely to have been explicitly trained to answer questions about this material. Smaller models are often quite effective at these sorts of RAG-style problems since they do not need to have internalised lots of facts and general information. In fact, in their technical report, the authors of Phi-3 mini call out the fact that their model can't retain factual information but could be augmented with search to produce reasonable results. This seemed like a perfect opportunity to test it out.
How The Task Works
There are 4 types of question in task B. Factoid, Yes/No, List and Summary. However, since summary is quite tricky to measure, it is not part of the BioASQ leaderboard. We also chose to omit summary from our tests.
Each question is provided along with a set of snippets. These are full sentences or paragraphs that have been pre-extracted from scientific papers. Incidentally, that activity is BioASQ Task A and it requires a lot more moving parts since there's retrieval involved too. In Task B we are concerned with existing sets of snippets and questions only.
In each case the model is required to respond with a short and precice exact answer to the question. The model may optionally also provide an ideal answer which provides some rationale for that answer. The ideal answer may provide useful context for the user but is not formally evaluated as part of BioASQ.
Yes/No questions require an exact answer of just "yes" or "no". For List questions, we are looking for a list of named entities (for example symptoms or types of microbe). For factoid we are typically looking for a single named entity. Models are allowed to respond to factoids with multiple answer. Therefore, factoids answers are scored by how closely to the top of the list the "correct" answer is ranked.
The Figure from the Hseuh et al 2023 Paper below illustrates this quite well:
![Examples of different question types. Full transcriptions of each are:
<p>Yes/No
Question: Proteomic analyses need prior knowledge of the organism complete genome. Is the complete genome of the bacteria of the genus Arthrobacter available?
Exact Answer: yes
Ideal Answer: Yes, the complete genome sequence of Arthrobacter (two strains) is deposited in GenBank.</p>
<p>List
Question: List Hemolytic Uremic Syndrome Triad.
Exact Answer: [anaemia, thrombocytopenia, renal failure]
Ideal Answer: Hemolytic uremic syndrome (HUS) is a clinical syndrome characterized by the triad of anaemia, thrombocytopenia, renal failure.</p>
<p>Factoid
Question: What enzyme is inhibited by Opicapone?
Exact Answer: [catechol-O-methyltransferase]
Ideal Answer: Opicapone is a novel catechol-O-methyltransferase (COMT) inhibitor to be used as adjunctive therapy in levodopa-treated patients with Parkinson’s disease</p>
<p>Summary
Question: What kind of affinity purification would you use in order to isolate soluble lysosomal proteins?
Ideal Answer: The rationale for purification of the soluble lysosomal proteins resides in their characteristic sugar, the mannose-6-phosphate (M6P), which allows an easy purification by affinity chromatography on immobilized M6P receptors." class=“wp-image-2510”/><figcaption class=](/media/image-8_eb742188.png) Figure 1 from the Hseuh et al 2023 Paper illustrates the different task types succinctly
Figure 1 from the Hseuh et al 2023 Paper illustrates the different task types succinctlyOur Setup
We wrote a python script that passes the question, context and guidance about the type of question to the model. We used a patched version of Ollama that allowed us to put restrictions on the shape of the model output. This allowed us to ensure responses were valid JSON in the same shape and structure as the BioASQ examples. These forced grammars saved us loads of time trying to coax JSON out of models in the structure we want. This is something that smaller models aren't great at. Sometimes models would still fail to give valid responses. For example, sometimes they get stuck in infinite loops spitting out brackets or newlines. We gave models up to 3 chances to produce a JSON response before a question is marked unanswerable and skipped.
Prompts
We used exactly the same prompts for all of the models which may have left room for further performance improvements. The exact prompts and grammar constraints that we used can be found here. Snippets are concatenated together with newlines in between them and provided as "context" in the prompt template.
We used the official BioASQ scoring tool to evaluate the responses and produce the results below. We evaluated our pipeline on the Task 11B Golden Enriched test set. You have to create a free account at bioasq to log in and download the data.
Models
We compared quantized versions of Phi and Llama with some other similarly sized models which perform well at benchmarks.
Note that although Phi is approx. half the size of the other models, the authors report competitive results against much larger models for a number of widely used benchmarks so it seems reasonable to compare it with these 7B and 8B models as oppose to only benchmarking against other 4B and smaller models.
Results
Yes/No Questions
The simplest type of BioASQ question is Yes/No. These results are measured with macro F1 to allow us to get a single metric across the performance at both "yes" and "no" questions.
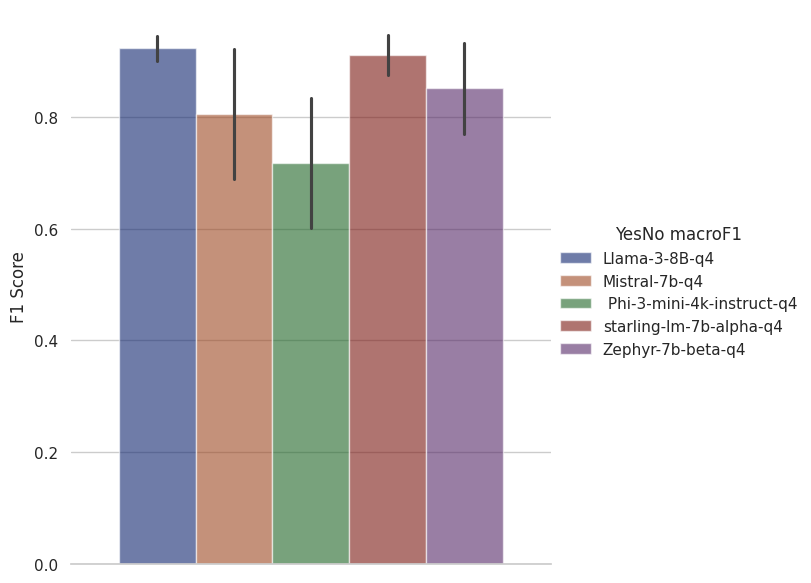 The best solutions to this task are coming in at 1.0 F1. Llama3 and Starling both achieve pretty close to perfect results here.
The best solutions to this task are coming in at 1.0 F1. Llama3 and Starling both achieve pretty close to perfect results here.
Factoid Questions
For factoid answers we measure responses in MRR since the model can return multiple possible answers. We are interested in how close the right answers are to the top of the list.
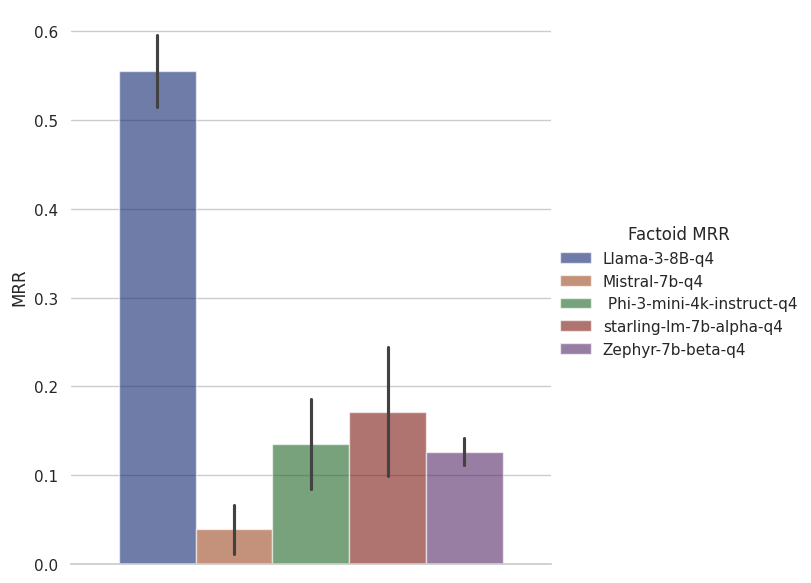 a GPT-4-based entrant, weighs in at 0.6316 MRR so it's pretty impressive that Llama 3 8B is providing results in the same ballpark as a model many times larger. For this one, Phi is in third place after Starling-LM 7B. Again, given that Phi is half the size of this model, it's quite impressive performance.
a GPT-4-based entrant, weighs in at 0.6316 MRR so it's pretty impressive that Llama 3 8B is providing results in the same ballpark as a model many times larger. For this one, Phi is in third place after Starling-LM 7B. Again, given that Phi is half the size of this model, it's quite impressive performance.
List Questions
We measure list questions in F1. A false positive is when something irrelevant is included in the answer and a false negative is when something relevant is missed from an answer. F1 gives us a single statistic that balances the two.
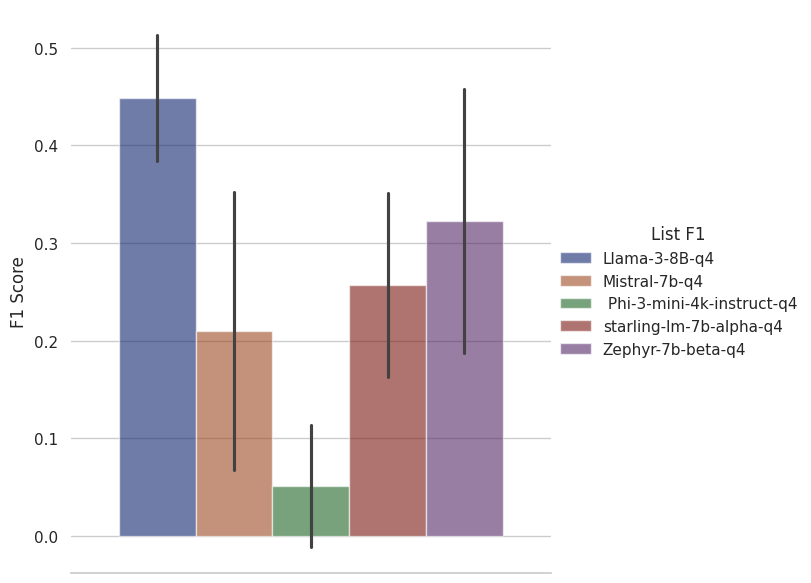 a GPT-4-based system, achieves an F1 of 0.72 so even Llama 3 8B leaves a relatively wide gap here. It would be worth testing the larger variants of Llama 3 to see how well they perform at this task and whether they are competitive with GPT-4.
a GPT-4-based system, achieves an F1 of 0.72 so even Llama 3 8B leaves a relatively wide gap here. It would be worth testing the larger variants of Llama 3 to see how well they perform at this task and whether they are competitive with GPT-4.
Discussion and Conclusion
Llama3
We've seen that Llama 3 8B and, to a lesser extent, Phi 3 Mini, are able to generalise reasonably well to a reading comprehension task in a field that wasn't a primary concern for either se of model authors. This isn't conclusive evidence for or against the general performance of these models on unseen tasks. However, it is certainly an interesting data point that shows that, particularly Llama 3 really is competitive with much larger models at this task. I wonder if that's because it was trained on such a large corpus which may have included some biomedical content as part of its training corpus.
Phi
I'm reluctant to too-harshly critique Phi's reasoning and reading comprehension ability since there's a good chance that it was disadvantaged by our test setup and the forced JSON structure, particularly for the list task. However, the weaker performance at the yes/no questions may be a hint that it isn't quite as good at generalised reading comprehension as the competing larger models.
We know that Phi3, like it's predecessor was trained on data that "consists of heavily filtered web data (according to the “educational level”) from various open internet sources, as well as synthetic LLM-generated data." However, we don't know specifically what was included or excluded. If Llama 3 went for "cast the net wide" approach to data collection, it's likely that the latter model may have been exposed to more biomedical content "by chance" and thus be better at reasoning about concepts that perhaps Phi had never seen before.
I do want to again call out that Phi is approximately half the size of the next biggest model in our benchmark so it's performance is quite impressive in that light.
Further Experiments
Model Size
I won't conjecture about whether 3.8B parameters is "too small" to generalise given the issues mentioned above but I'd love to see some more tests of this in future. Do the larger variants of Phi (trained on the same data but simpliy with more parameters) suffer from the same issues?
Model Fine Tuning
The models that I've been testing are small enough that they can be fine-tuned on specific problems on a consumer-grade gaming GPU for very little cost. It seems entirely plausible to me that by fine-tuning these models on biomedical text ands historical BioASQ training sets their performance could be improved even more significantly. The challenge would be in finding the right mix of data.
Better Prompts
We did not spend a lot of time attempting to build effective prompts during this experiment. It may be that performance was left on the table because of this oversight. Smaller models are often quite fussy about prompts. It might be interesting to use a prompt optimisation framework like DSPy to be more systematic about better prompts.
Other Tasks
I tried these models on BioAsq but this is lightyears away from conclusive evidence for whether or not these new-generation small models can generalise well. It's simply a test of whether they can do biology. It will be very interesting to try other novel tasks and see how well they perform. Watch this space!
Replies & Web Activities
If you would like to comment or reply then toot me or bluesky me about this url, or send me a webmention